
- Read more about Fourier Ptychography Microscopy With Integrated Positional Misalignment Correction
- Log in to post comments
Fourier Ptychography Microscopy enables reconstructing both intensity and phase high-resolution wide-field images from multiple captures under varying illumination directions. The capture process is classically modeled using a neural network. The reconstructed object is iteratively optimized by gradient descent so the network output matches the captures. Although, this process hinges on a precise estimation of the system geometry.
- Categories:
 49 Views
49 Views- Read more about Transforming Tabular Data For Multi-modality: Enhancing Breast Cancer Metastasis Prediction Through Data Conversion
- Log in to post comments
Breast cancer metastasis prediction plays a key role in
clinical decision-making and secondary analysis. Traditionally,
metastasis classification models have been developed
using structured tabular clinical data, but these approaches
may result in data loss and lack of contextual information. A
multi-modal approach is presented in this article for predicting
breast cancer metastasis by converting structured clinical
data into unstructured text, which provides more contextual
information, and then converting that text into histopathology
- Categories:
 31 Views
31 Views
- Read more about A registration error estimation framework for correlative imaging
- Log in to post comments
Correlative imaging workflows are now widely used in bio-imaging and aims to image the same sample using at least two different and complementary imaging modalities. Part of the workflow relies on finding the transformation linking a source image to a target image. We are specifically interested in the estimation of registration error in point-based registration.
poster.pdf
- Categories:
 74 Views
74 Views- Read more about Super-Resolved Imaging of Early-Stage Dynamics in The Immune Response
- Log in to post comments
The use of photo-activated fluorescent molecules to create long sequences of low-density, diffraction-limited images gives us the ability to achieve highly-precise molecule localizations. However, this methodology requires lengthy imaging times, resulting in poor temporal resolution. This is particularly problematic when dynamic interactions of live cells on short time scales are of interest. We consider the problem of shortening dramatically the acquisition times in super-resolution microscopy down to seconds, in order to image the cellular dynamics during T-cell activation.
- Categories:
 69 Views
69 Views
Segmentation of biological images is a challenging task, due to non convex shapes, intensity inhomogeneity and clustered cells. To address these issues, a new algorithm is proposed based on the B-spline level set method. The implicit function of the level set is modelled as a continuous parametric function represented with the B-spline basis. It is different from the discrete formulation associated with conventional level set. In this paper the proposed framework takes into account properties of biological images.
- Categories:
 35 Views
35 ViewsHyperspectral (HS) imaging retrieves information from data obtained across a wide spectral range of spectral channels. The object to reconstruct is a 3D cube, where two coordinates are spatial and the third one is spectral. We assume that this cube is complex-valued, i.e. characterized spatially frequency varying amplitude and phase. The observations are squared magnitudes measured as intensities summarized over the spectrum. The HS phase retrieval problem is formulated as a reconstruction of the HS complex-valued object cube from Gaussian noisy intensity observations.
- Categories:
 50 Views
50 Views
- Read more about Signal Sensing and Reconstruction Paradigms for a Novel Multi-source Static Computed Tomography System
- Log in to post comments
Conventional Computed Tomography (CT) systems use a single X-ray source and an arc of detectors mounted on a rotating gantry to acquire a set of projection data. Novel CT systems are now being pioneered in which a complete ring of distributed X-ray sources and detectors are electronically turned on and off, without any mechanical motion, to acquire a set of projections for tomographic reconstruction. This paper discusses new sensing and reconstruction paradigms enabled by this new CT architecture.
- Categories:
 82 Views
82 Views
- Read more about LEARNING SPATIALLY-CORRELATED TEMPORAL DICTIONARIES FOR CALCIUM IMAGING
- Log in to post comments
Calcium imaging has become a fundamental neural imaging technique, aiming to recover the individual activity of hundreds of neurons in a cortical region. Current methods (mostly matrix factorization) are aimed at detecting neurons in the field-of-view and then inferring the corresponding time-traces. In this paper, we reverse the modeling and instead aim to minimize the spatial inference, while focusing on finding the set of temporal traces present in the data.
ICASSP.pdf
- Categories:
 9 Views
9 Views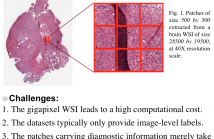
- Read more about WHOLE SLIDE IMAGE CLASSIFICATION VIA ITERATIVE PATCH LABELLING
- Log in to post comments
Brain tumor can be a fatal disease in the world. With the aim of improving survival rates, many computerized algorithms have been proposed to assist the pathologists to make a diagnosis, using Whole Slide Pathology Images (WSI). Most methods focus on performing patch-level classification and aggregating the patch-level results to obtain the image classification. Since not all patches carry diagnostic information, it is thus important for our algorithm to recognize discriminative and non-discriminative patches.
- Categories:
 184 Views
184 Views