
Optimal power flow is a critical optimization problem that allocates power to the generators in order to satisfy the demand at a minimum cost. This is a non-convex problem shown to be NP-hard. We use a graph neural network to learn a nonlinear function between the power demanded and the corresponding allocation. We learn the solution in an unsupervised manner, minimizing the cost directly. To consider the power system constraints, we propose a novel barrier method that is differentiable and works on initially infeasible points.
- Categories:
 59 Views
59 Views
- Read more about A graph-prediction-based approach for debiasing underreported data
- Log in to post comments
We present a novel Graph-based debiasing Algorithm for Underreported Data (GRAUD) aiming at an efficient joint estimation of event counts and discovery probabilities across spatial or graphical structures. This innovative method provides a solution to problems seen in fields such as policing data and COVID-19 data analysis. Our approach avoids the need for strong priors typically associated with Bayesian frameworks. By leveraging the graph structures on unknown variables n and p, our method debiases the under-report data and estimates the discovery probability at the same time.
- Categories:
 16 Views
16 Views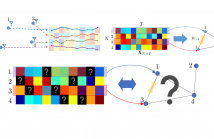
- Read more about Joint Signal Recovery and Graph Learning from Incomplete Time-Series
- Log in to post comments
Learning a graph from data is the key to taking advantage of graph signal processing tools. Most of the conventional algorithms for graph learning require complete data statistics, which might not be available in some scenarios. In this work, we aim to learn a graph from incomplete time-series observations. From another viewpoint, we consider the problem of semi-blind recovery of time-varying graph signals where the underlying graph model is unknown.
- Categories:
 69 Views
69 Views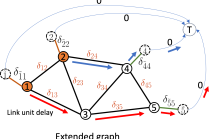
- Read more about Congestion-aware Distributed Task Offloading in Wireless Multi-hop Networks using Graph Neural Networks
- Log in to post comments
Computational offloading has become an enabling component for edge intelligence in mobile and smart devices. Existing offloading schemes mainly focus on mobile devices and servers, while ignoring the potential network congestion caused by tasks from multiple mobile devices, especially in wireless multi-hop networks. To fill this gap, we propose a low-overhead, congestion-aware distributed task offloading scheme by augmenting a distributed greedy framework with graph-based machine learning.
- Categories:
 18 Views
18 Views
- Read more about Reducing the Communication and Computational Cost of Random Fourier Features Kernel LMS in Diffusion Networks
- Log in to post comments
Diffusion kernel algorithms are interesting tools for distributed nonlinear estimation. However, for the sake of feasibility, it is essential in practice to restrict their computational cost and the number of communications. In this paper, we propose a censoring algorithm for adaptive kernel diffusion networks based on random Fourier features that locally adapts the number of nodes censored according to the estimation error.
- Categories:
 23 Views
23 Views
In this paper, we revisit the popular affinity matrix based on the anchor graph and point out that the spectral embedding obtained using symmetric normalized Laplacian is only a side view of the bipartite structure. Based on the analysis, we propose Fast Spectral Clustering based on the Random Walk Laplacian (FRWL) method to explicitly balance the popularity of anchors and the independence of data points, which is especially important for clustering of boundary points.
- Categories:
 54 Views
54 Views
- Read more about Generalized Kernel-Based Dynamic Mode Decomposition
- 2 comments
- Log in to post comments
manuscript.pdf
- Categories:
 25 Views
25 Views
- Read more about Kernel Node Embeddings
- Log in to post comments
Learning representations of nodes in a low dimensional space is a crucial task with many interesting applications in network analysis, including link prediction and node classification. Two popular approaches for this problem include matrix factorization and random walk-based models. In this paper, we aim to bring together the best of both worlds, towards learning latent node representations. In particular, we propose a weighted matrix factorization model which encodes random walk-based information about the nodes of the graph.
- Categories:
 34 Views
34 Views
- Read more about Graph Filtering with Multiple Shift Matrices
- Log in to post comments
- Categories:
 35 Views
35 Views