
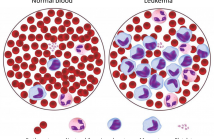
- Read more about Acute Lymphoblastic Leukemia detection based on adaptive unsharpening and Deep Learning
- Log in to post comments
Computer Aided Diagnosis (CAD) systems are increasingly utilizing image analysis and Deep Learning (DL) techniques, due to their high accuracy in several medical imaging fields, including the detection of Acute Lymphoblastic (or Lymphocytic) Leukemia (ALL) from peripheral blood samples. However, no method in the literature has specifically analyzed the focus quality of ALL images or proposed a technique for sharpening the samples in an adaptive way for the purpose of classification.
- Categories:
 51 Views
51 Views
- Read more about Instance segmentation with the number of clusters incorporated in embedding learning
- Log in to post comments
Poster.pdf
- Categories:
 4 Views
4 Views
- Read more about SEMI-SUPERVISED SKIN LESION SEGMENTATION WITH LEARNING MODEL CONFIDENCE
- Log in to post comments
poster-xzq.pdf
- Categories:
 4 Views
4 Views
- Read more about SEMI-SUPERVISED SKIN LESION SEGMENTATION WITH LEARNING MODEL CONFIDENCE
- Log in to post comments
poster-xzq.pdf
- Categories:
 17 Views
17 Views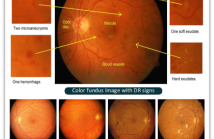
- Read more about SEA-NET: SQUEEZE-AND-EXCITATION ATTENTION NET FOR DIABETIC RETINOPATHY GRADING
- Log in to post comments
Diabetes is one of the most common disease in individuals. Diabetic retinopathy (DR) is a complication of diabetes, which could lead to blindness. Automatic DR grading based on retinal images provides a great diagnostic and prognostic value for treatment planning. However, the subtle differences among severity levels make it difficult to capture important features using conventional methods.
- Categories:
 420 Views
420 Views
- Read more about An enhanced deep learning architecture for classification of tuberculosis types from CT lung images
- Log in to post comments
- Categories:
 31 Views
31 Views
- Read more about A segmentation based deep learning framework for multimodal retinal image registration
- Log in to post comments
Multimodal image registration plays an important role in diagnosing and treating ophthalmologic diseases. In this paper, a deep learning framework for multimodal retinal image registration is proposed. The framework consists of a segmentation network, feature detection and description network, and an outlier rejection network, which focuses only on the globally coarse alignment step using the perspective transformation.
- Categories:
 49 Views
49 Views